-Search query
-Search result
Showing all 43 items for (author: trono & d)

EMDB-17819: 
XBB 1.0 RBD bound to P4J15 (Local)
Method: single particle / : Duhoo Y, Lau K
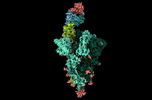
EMDB-17849: 
XBB 1.0 RBD bound to P4J15 (Global)
Method: single particle / : Duhoo Y, Lau K

EMDB-17850: 
SARS-CoV-2 XBB 1.0 closed conformation.
Method: single particle / : Duhoo Y, Lau K
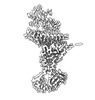
EMDB-15592: 
BA.4/5 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

PDB-8aqw: 
BA.4/5 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-14922: 
cryo-EM structure of omicron spike in complex with de novo designed binder, full map
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zrv: 
cryo-EM structure of omicron spike in complex with de novo designed binder, full map
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

EMDB-14930: 
cryo-EM structure of omicron spike in complex with de novo designed binder, local
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

EMDB-14947: 
cryo-EM structure of D614 spike in complex with de novo designed binder, full and local maps(addition)
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zsd: 
cryo-EM structure of omicron spike in complex with de novo designed binder, local
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC

PDB-7zss: 
cryo-EM structure of D614 spike in complex with de novo designed binder
Method: single particle / : Pablo G, Sarah W, Alexandra VH, Anthony M, Andreas S, Zander H, Dongchun N, Shuguang T, Freyr S, Casper G, Priscilla T, Alexandra T, Stephane R, Sandrine G, Jane M, Aaron P, Zepeng X, Yan C, Pu H, George G, Elisa O, Beat F, Didier T, Henning S, Michael B, Bruno EC
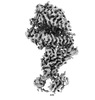
EMDB-15588: 
BA.4/5 SARS-CoV-2 Spike bound to human ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15589: 
Beta SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15590: 
BA.1 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E
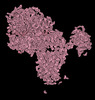
EMDB-15591: 
BA.2.12.1 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

PDB-8aqs: 
BA.4/5 SARS-CoV-2 Spike bound to human ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

PDB-8aqt: 
Beta SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

PDB-8aqu: 
BA.1 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

PDB-8aqv: 
BA.2.12.1 SARS-CoV-2 Spike bound to mouse ACE2 (local)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15541: 
Beta SARS-CoV-2 Spike bound to mouse ACE2 (two up, full)
Method: single particle / : Ni D, Lau K, Turelli P, Beckert B, Nazarov S, Myasnikov A, Trono D, Stahlberg H, Pojer F, Uchikawa E

EMDB-15580: 
BA.1 SARS-CoV-2 Spike bound to mouse ACE2 (two up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E
EMDB-15581: 
BA.1 SARS-CoV-2 Spike bound to mouse ACE2 (three up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15584: 
BA.2.12.1 SARS-CoV-2 Spike bound to mouse ACE2 (two up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15585: 
BA.2.12.1 SARS-CoV-2 Spike bound to mouse ACE2 (three up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15586: 
BA.4/5 SARS-CoV-2 Spike bound to mouse ACE2 (two up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-15587: 
BA.4/5 SARS-CoV-2 Spike bound to human ACE2 (three up, full)
Method: single particle / : Lau K, Ni D, Beckert B, Nazarov S, Myasnikov A, Pojer F, Stahlberg H, Uchikawa E

EMDB-14141: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - 3-P2G3 and 1-P5C3 Fabs (Global)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

EMDB-14142: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD up - 1-P2G3 and 1-P5C3 Fabs (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

EMDB-14143: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD down - 1-P2G3 Fab (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

PDB-7qti: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - 3-P2G3 and 1-P5C3 Fabs (Global)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

PDB-7qtj: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD up - 1-P2G3 and 1-P5C3 Fabs (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

PDB-7qtk: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD down - 1-P2G3 Fab (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Fenwick C, Perez L, Pojer F, Stahlberg H, Pantaleo G, Trono D

EMDB-14087: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD and NTD (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Beckert B, Nazarov S, Pojer F, Myasnikov A, Stahlberg H, Trono D

PDB-7qo9: 
SARS-CoV-2 S Omicron Spike B.1.1.529 - RBD and NTD (Local)
Method: single particle / : Ni D, Lau K, Turelli P, Beckert B, Nazarov S, Pojer F, Myasnikov A, Stahlberg H, Trono D

EMDB-14086: 
SARS-CoV-2 S Omicron Spike B.1.1.529
Method: single particle / : Ni D, Lau K, Turelli P, Beckert B, Nazarov S, Pojer F, Myasnikov A, Stahlberg H, Trono D

PDB-7qo7: 
SARS-CoV-2 S Omicron Spike B.1.1.529
Method: single particle / : Ni D, Lau K, Turelli P, Beckert B, Nazarov S, Pojer F, Myasnikov A, Stahlberg H, Trono D

EMDB-13190: 
P5C3 is a potent fab neutralizer
Method: single particle / : perez L

EMDB-13265: 
the local resolution of Fab p5c3.
Method: single particle / : perez L

EMDB-13415: 
MaP OF P5C3RBD Interface
Method: single particle / : Perez L
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search







 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
